Quantifying nuclei
The Problem
Immunohistochemistry is a high-specifity method that uses tagged antibodies to identify proteins in tissue. This method enables the use of fluorescent markers that glow when bound to the target protein and allow researchers to visualize the expression of the protein in the given tissue. Despite the popularity of this method, tools for quantifying structures remain largely inaccessible and researchers typically quantify the stained structures manually (loose estimation is much more common than anyone will readily admit), which is a burden on both the researcher’s time as well as the integrity of their data. Since you can stain for up to three proteins (“red”, “green”, “blue”), this would typically entail 3x the work load!
The Solution
To avoid having to do this task for many hundreds of images (the number of hours would have been incalculable), I built a system that would automatically identify and count structures from images of our stained tissue. The user simple inputs an area constraint (“the structure lies roughly within a range of A to B pixels^2”) and lets the computer do the rest! Let’s look at an example to understand how the program works:
1) Start with an image
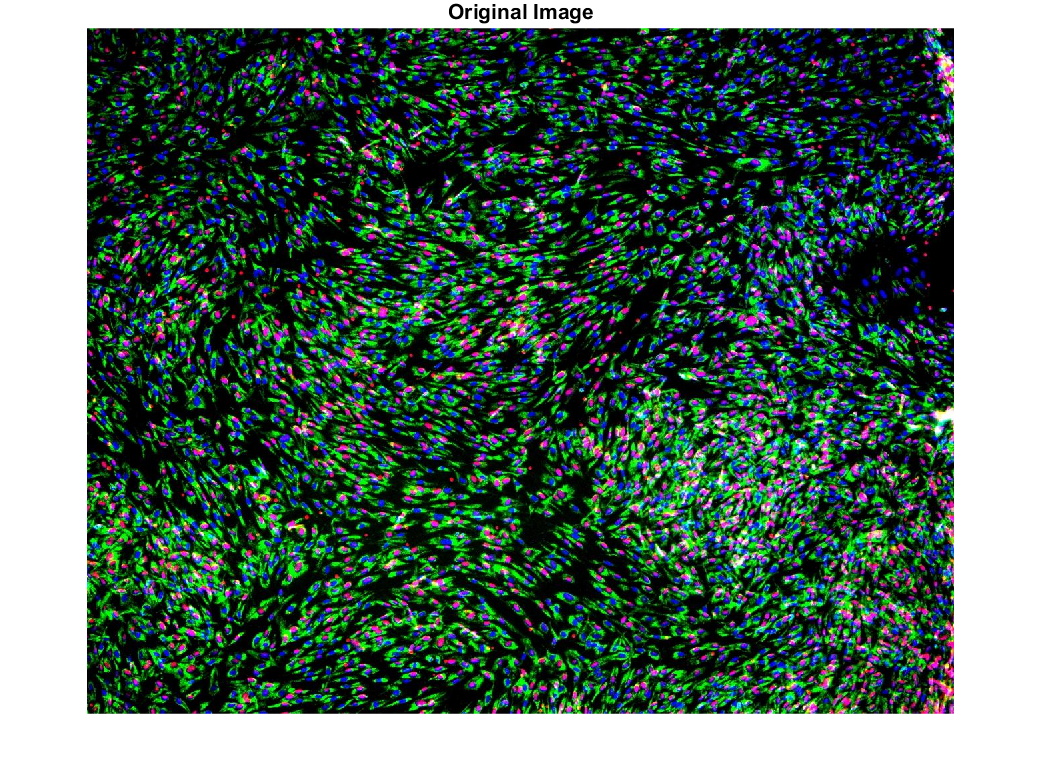
2) Split into color channels
3) Binarize the image and sharpen borders
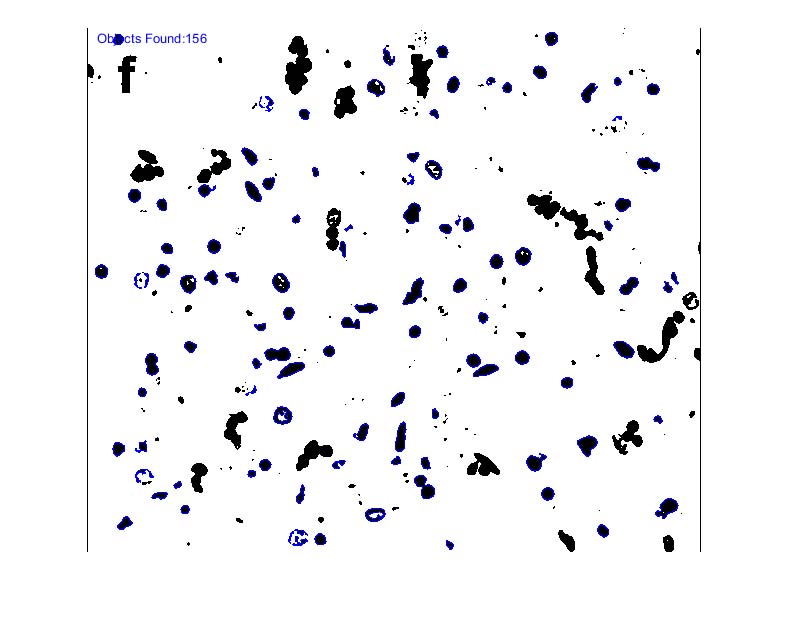
4) Identify objects
5) Count objects
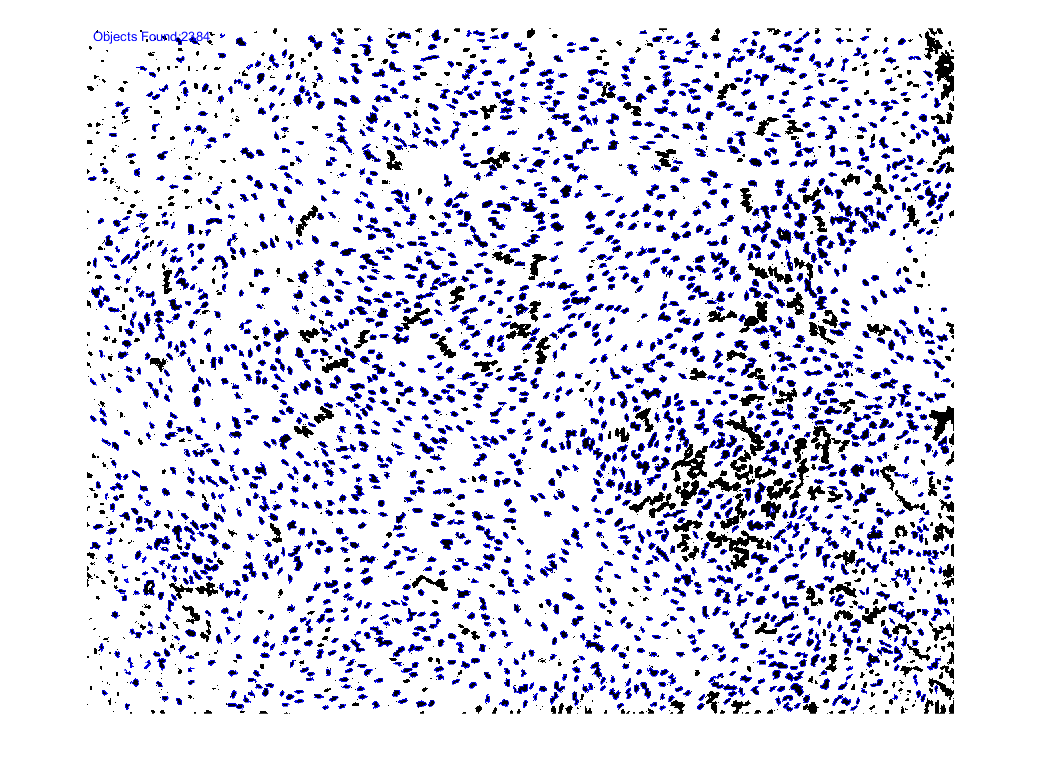

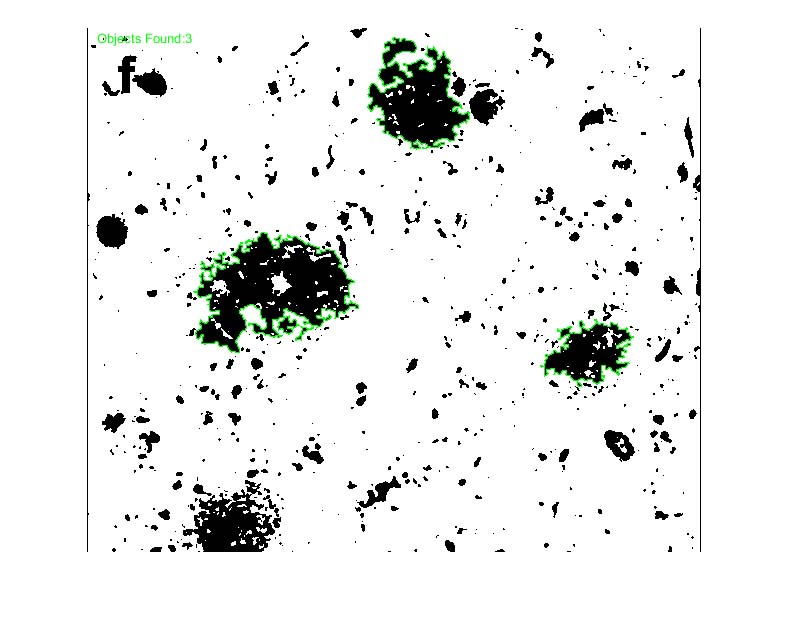
Note: you may notice that there are several objects that don’t appear to be included in the total count (no blue boundary around them… These objects are layered on top of each other (“occluded”) and as a result, the object detection algorithm sees them as one object. Since we set an initial constraint on the area of our desired object, we can get the rough number of objects in these clusters by taking the area of objects above our upper threshold (the area of these clusters) and dividing that by the average area of the objects within our range (objects identified in blue).
Now another example, this time with morphologically weird objects (clear-cut circles, like the nuclei above, are easiest)
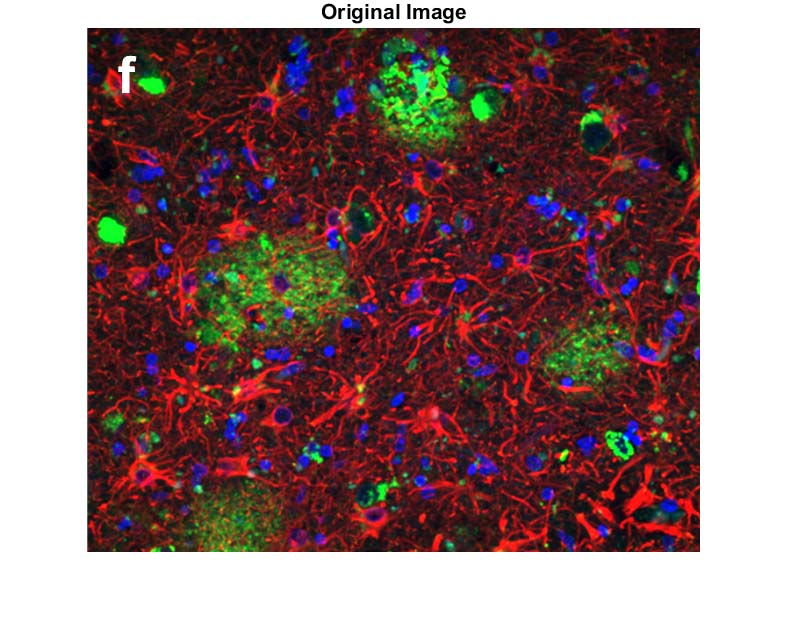
Now we split the image into each of the RGB channels: